-Search query
-Search result
Showing 1 - 50 of 177 items for (author: gonzalez & b)

EMDB-17626: 
E. coli RNA polymerase paused at ops site
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17632: 
fully recruited RfaH bound to E. coli transcription complex paused at ops site (alternative state of RfaH)
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17646: 
transcription complex paused at ops site and bound to autoinhibited RfaH, not fully complementary scaffold
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17647: 
fully recruited RfaH bound to E. coli transcription complex paused at ops site (not fully complementary scaffold; alternative state of RfaH)
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17657: 
E. coli RNA polymerase paused at ops site (non-complementary scaffold)
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17668: 
E. coli transcription complex paused at ops site with fully recruited RfaH
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17679: 
autoinhibited RfaH bound to E. coli transcription complex paused at ops site (encounter complex)
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17681: 
backtracked E. coli transcription complex paused at ops site and bound to RfaH
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17685: 
E. coli transcription complex paused at ops site and bound to RfaH and NusA
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-17686: 
fully recruited RfaH bound to E. coli transcription complex paused at ops site (not complementary scaffold)
Method: single particle / : Zuber PK, Said N, Hilal T, Loll B, Wahl MC, Knauer SH

EMDB-19194: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #2
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-19346: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #1
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-40223: 
Cryo-ET reconstruction of WT HSV-1 NEC coat
Method: subtomogram averaging / : Wang H, Draganova EB

EMDB-40224: 
Cryo-ET reconstruction of HSV-1 DN/SUP mutant NEC coat
Method: subtomogram averaging / : Wang H, Draganova EB

EMDB-40225: 
Cryo-ET reconstruction of HSV-1 SUP mutant NEC coat
Method: subtomogram averaging / : Wang H, Draganova EB
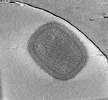
EMDB-18916: 
Cryotomogram of mature Vaccinia virus (WR) virion
Method: electron tomography / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-18917: 
Subtomogram average of the Vaccinia virus (WR) portal complex in mature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-18918: 
Subtomogram average of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

PDB-8r5i: 
In situ structure of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions by flexible fitting into a cryoET map
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-41171: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L

EMDB-41172: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L
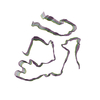
PDB-8tdn: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L
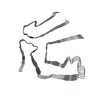
PDB-8tdo: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L

EMDB-16778: 
Type six secretion system exported effector 5 (Tse5)
Method: single particle / : Gonzalez-Magana A, Tascon I, Ubarretxena-Belandia I, Albesa-Jove D

PDB-8cp6: 
Type six secretion system exported effector 5 (Tse5)
Method: single particle / : Gonzalez-Magana A, Tascon I, Ubarretxena-Belandia I, Albesa-Jove D

EMDB-16024: 
SARS-CoV-2 S protein in complex with pT1644 Fab
Method: single particle / : Stroeh L, Hansen G, Vollmer B, Krey T, Benecke T, Gruenewald K

EMDB-16026: 
SARS-CoV-2 S protein in complex with pT1696 Fab
Method: single particle / : Hansen G, Benecke T, Vollmer B, Gruenewald K, Krey T

EMDB-27781: 
Cryo-EM structure of 227 Fab in complex with (NPNA)8 peptide
Method: single particle / : Martin GM, Ward AB

EMDB-27784: 
Cryo-EM structure of 239 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27785: 
Cryo-EM structure of 311 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27786: 
Cryo-EM structure of 334 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27787: 
Cryo-EM structure of 337 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27788: 
Cryo-EM structure of 356 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27789: 
Cryo-EM structure of 364 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dyt: 
Cryo-EM structure of 227 Fab in complex with (NPNA)8 peptide
Method: single particle / : Martin GM, Ward AB

PDB-8dyw: 
Cryo-EM structure of 239 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dyx: 
Cryo-EM structure of 311 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dyy: 
Cryo-EM structure of 334 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dz3: 
Cryo-EM structure of 337 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dz4: 
Cryo-EM structure of 356 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dz5: 
Cryo-EM structure of 364 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-15694: 
Human adenovirus type 5 lacking core protein V
Method: single particle / : Hernando-Perez M, San Martin C, Martin-Gonzalez N, Gomez-Gonzalez A, Bauer M, Greber UF, de Pablo PJ

EMDB-40455: 
Cryo-EM structure of Karyopherin-beta2 bound to HNRNPH2 PY-NLS
Method: single particle / : Gonzalez A, Fung HYJ, Chook YM

PDB-8sgh: 
Cryo-EM structure of Karyopherin-beta2 bound to HNRNPH2 PY-NLS
Method: single particle / : Gonzalez A, Fung HYJ, Chook YM

EMDB-17541: 
Cryo-EM density map of UBR5 in complex with MCRS1
Method: single particle / : Schmitt S, Aguirre JD, Kempf G, Kater L, Thoma NH

EMDB-26685: 
Cardiac amyloid fibrils extracted from a variant ATTR I84S amyloidosis patient
Method: helical / : Nguyen BA, Saelices L

EMDB-27323: 
Cardiac amyloid fibrils extracted from a variant ATTR I84S amyloidosis patient (2).
Method: helical / : Nguyen BA, Saelices L

EMDB-29035: 
Prefusion-stabilized SARS-CoV-2 spike protein
Method: single particle / : Gonzalez KJ, Mousa JJ, Strauch EM

PDB-8fez: 
Prefusion-stabilized SARS-CoV-2 spike protein
Method: single particle / : Gonzalez KJ, Mousa JJ, Strauch EM

EMDB-15596: 
In situ subtomogram average of Vaccinia virus (WR) palisade, all virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
